| Clin Mol Hepatol > Volume 27(2); 2021 > Article |
|
ABSTRACT
Background/Aims
Methods
Results
ACKNOWLEDGMENTS
FOOTNOTES
SUPPLEMENTAL MATERIAL
Supplementary┬ĀData┬Ā1.
Supplementary┬ĀData┬Ā2.
Figure┬Ā1.
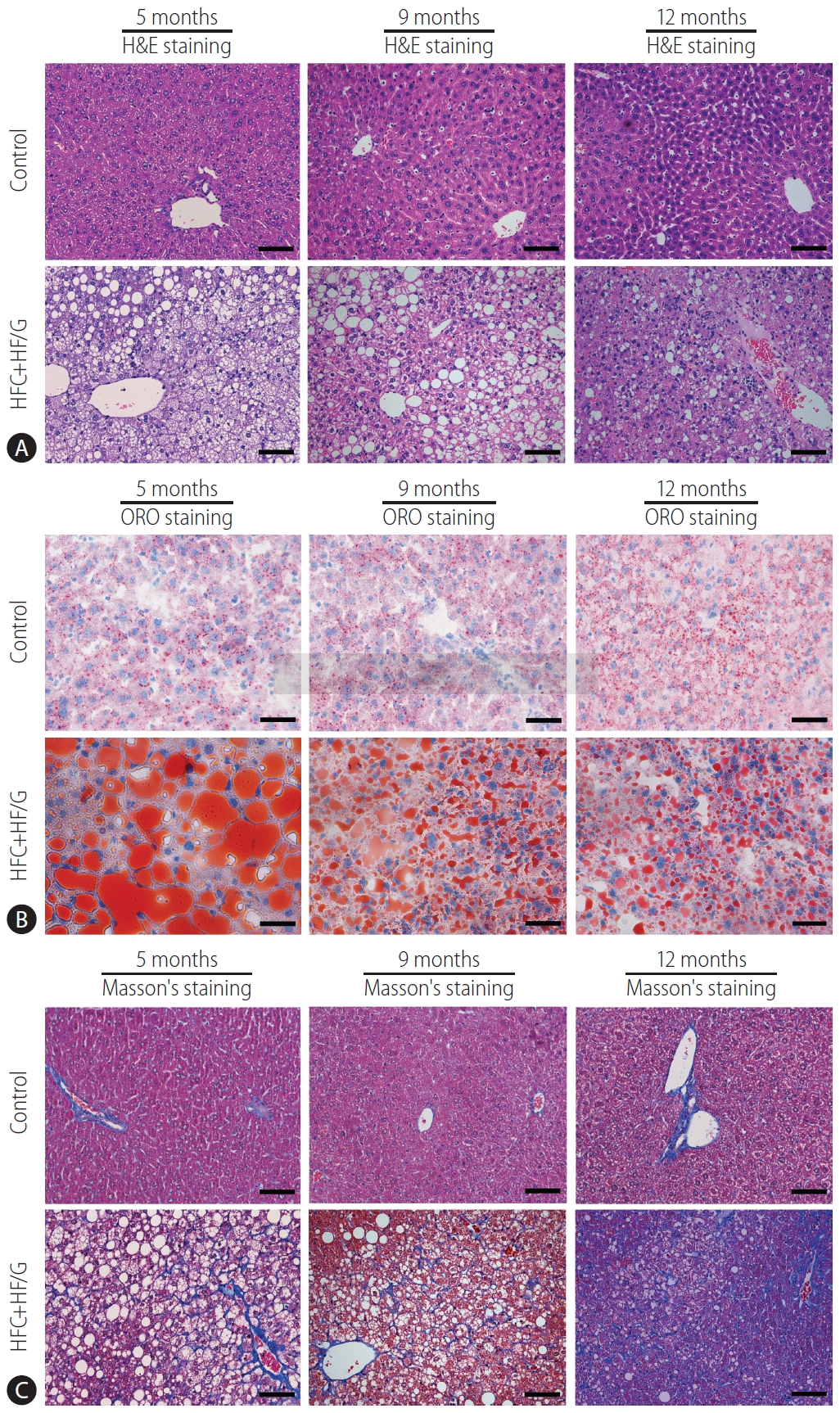
Figure┬Ā2.
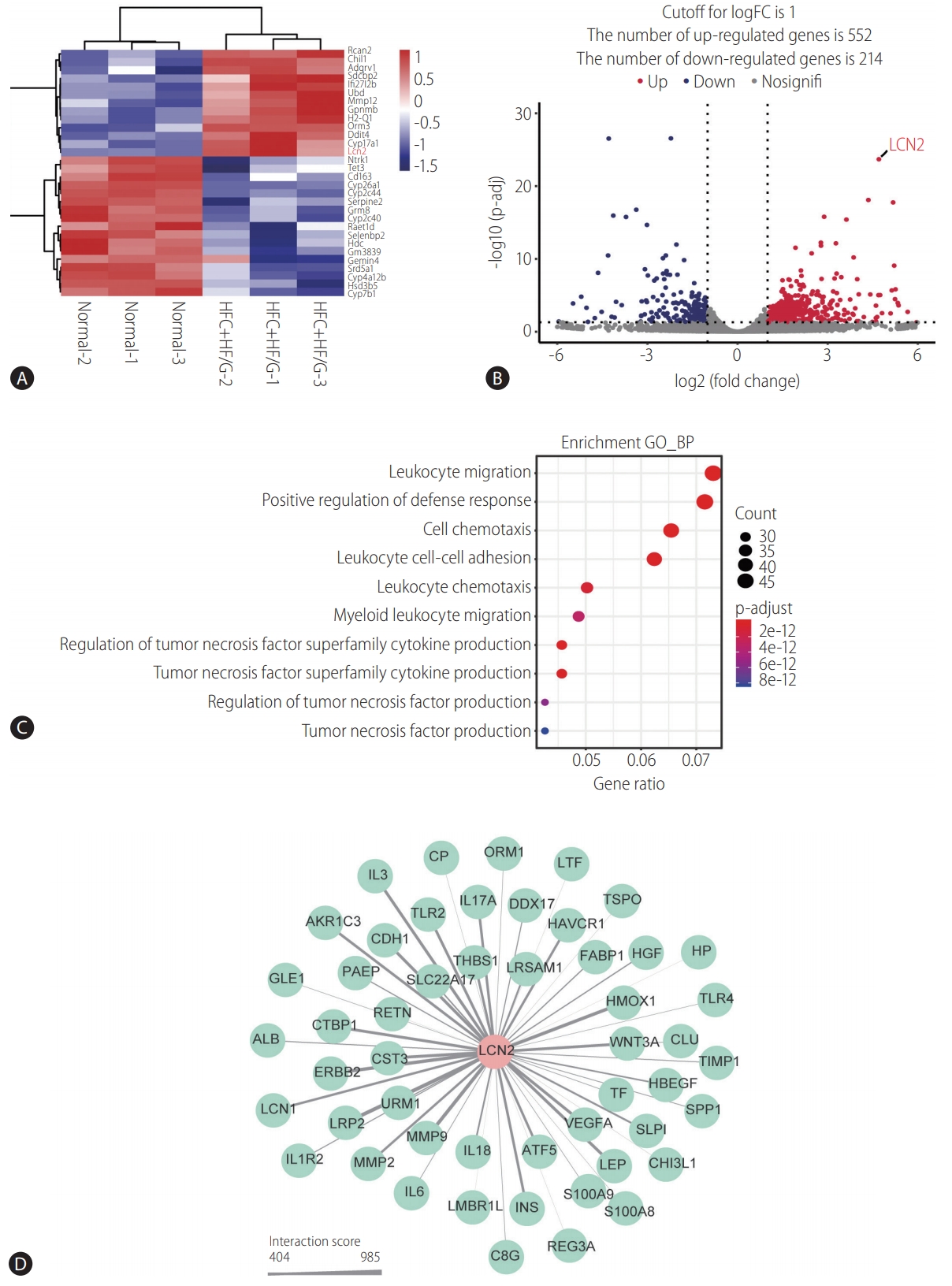
Figure┬Ā3.
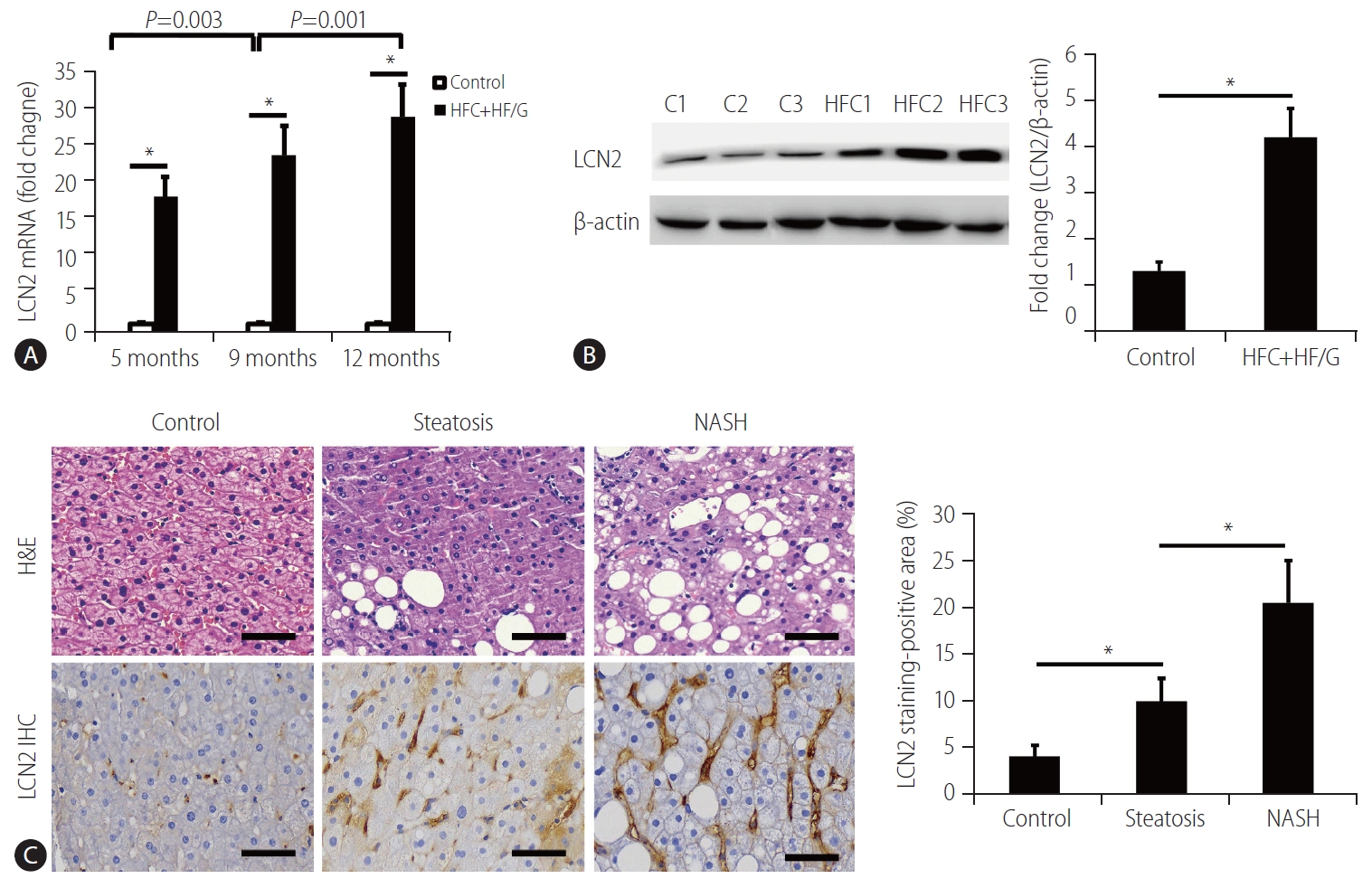
Figure┬Ā4.

Figure┬Ā5.

Table┬Ā1.
| Factor | NASH group (n=277) | Steatosis group (n=83) | Control group (n=40) | F/Žć2 | P-value |
|---|---|---|---|---|---|
| Age (years) | 39.92┬▒11.99* | 45.07┬▒12.09 | 41.43┬▒11.48 | 5.936 | 0.003 |
| Height | 168.06┬▒8.16 | 166.78┬▒7.80 | 168.25┬▒6.84 | 0.885 | 0.413 |
| Weight | 76.49┬▒14.07*,ŌĆĀ | 70.18┬▒10.12ŌĆĀ | 61.35┬▒11.00 | 27.227 | <0.001 |
| BMI (kg/m2) | 26.95┬▒3.65*,ŌĆĀ | 25.16┬▒2.59ŌĆĀ | 21.50┬▒2.51 | 49.593 | <0.001 |
| Sex | 4.199 | 0.123 | |||
| ŌĆāFemale | 73 (26.4) | 18 (21.7) | 16 (40.0) | ||
| ŌĆāMale | 204 (73.6) | 65 (78.9) | 24 (60.0) | ||
| Blood LCN2 | 85.20 (64.24, 107.25)*,ŌĆĀ | 66.25(47.18, 88.66)ŌĆĀ | 32.32 (24.80, 41.00) | 98.909 | <0.001 |
| Total bilirubin | 13.0 (10.0, 17.0) | 12.0 (10.0, 15.0) | 12.0 (9.0, 14.0) | 4.734 | 0.094 |
| Direct bilirubin | 5.0 (4.0, 6.0) | 4.0 (4.0, 6.0) | 5.0 (4.0, 6.0) | 0.646 | 0.724 |
| Indirect bilirubin | 8.0 (6.0, 11.0)ŌĆĀ | 7.0 (6.0, 9.0) | 7.0 (4.3, 10.0) | 6.926 | 0.031 |
| Total cholesterol | 5.09 (4.36, 5.89)*,ŌĆĀ | 4.67 (3.93, 5.41)ŌĆĀ | 4.17 (3.87, 4.43) | 36.283 | <0.001 |
| Triglyceride | 1.92 (1.45, 2.81)*,ŌĆĀ | 1.72 (1.22, 2.33)ŌĆĀ | 1.24 (0.89, 1.40) | 52.112 | <0.001 |
| HDL cholesterol | 0.99 (0.87, 1.16)ŌĆĀ | 1.00 (0.87, 1.12)ŌĆĀ | 1.16 (0.99, 1.41) | 14.031 | 0.001 |
| LDL cholesterol | 3.16 (2.43, 3.75)ŌĆĀ | 2.83 (2.38, 3.38)ŌĆĀ | 2.31 (1.86, 2.72) | 35.46 | <0.001 |
| Total protein | 78.0 (74.2, 80.9)*,ŌĆĀ | 76.0 (72.6, 78.9) | 76.6 (72.0, 80.4) | 11.227 | 0.004 |
| Albumin | 46.8 (44.4, 49.0)* | 45.1 (42.6, 47.6) | 46.1 (43.4, 48.3) | 12.679 | 0.002 |
| Alanine aminotransferase | 60.0(36.5, 107.5)*,ŌĆĀ | 40.0 (23, 70.0)ŌĆĀ | 23.0 (17.3, 26.8) | 81.829 | <0.001 |
| Aspartate aminotransferase | 38.0 (27.0, 61.0)*,ŌĆĀ | 30.0 (23.0, 41.0)ŌĆĀ | 22.0 (20.0, 25.0) | 68.2 | <0.001 |
| Gluten and aspartate ratio | 1.54 (1.20, 1.92)*,ŌĆĀ | 1.27 (0.90, 1.60)ŌĆĀ | 1.04 (0.88, 1.04) | 53.983 | <0.001 |
| Alkaline phosphatase | 81.0 (69.0, 99.0)ŌĆĀ | 77.0 (66.0, 96.0) | 79.0 (63.0, 92.5) | 1.514 | 0.469 |
| Glutamyl transferase | 53.0 (34.0, 86.5)ŌĆĀ | 44.0 (26.0, 80.0)ŌĆĀ | 22.0 (16.0, 28.0) | 69.365 | <0.001 |
| Glucose | 5.2 (4.8, 6.0)ŌĆĀ | 5.2 (4.7, 6.4)ŌĆĀ | 4.7 (4.3, 5.1) | 26.752 | <0.001 |
| Urea nitrogen | 4.6 (3.9, 5.4)*,ŌĆĀ | 4.9 (4.2, 6.0) | 5.2 (4.4, 5.8) | 10.391 | 0.006 |
| Creatinine | 67.0 (56.5, 75.5)* | 71.0 (63.0, 79.0)ŌĆĀ | 64.5 (56.3, 73.5) | 7.9 | 0.019 |
| Fasting insulin | 111.0 (78.6, 156.3)*,ŌĆĀ | 78.3 (52.6, 120.4)ŌĆĀ | 14.8 (12.3, 19.3) | 117.623 | <0.001 |
| Steatosis | 193.87 | <0.001 | |||
| ŌĆā<5% | 0 (0.0)ŌĆĀ | 0 (0.0)ŌĆĀ | 40 (100.0) | ||
| ŌĆā5ŌĆō33% | 91 (32.9)*,ŌĆĀ | 55 (66.3)ŌĆĀ | 0 (0.0) | ||
| ŌĆā34ŌĆō66% | 119 (43.0)* | 21(25.3) | 0 (0.0) | ||
| ŌĆā>66% | 67 (24.1)* | 7 (8.4) | 0 (0.0) | ||
| Balloon-like change | 400 | <0.001 | |||
| ŌĆāNon | 0 (0.0)*,ŌĆĀ | 83 (100.0) | 40 (100.0) | ||
| ŌĆāRare | 214 (77.3) | 0 (0.0) | 0 (0.0) | ||
| ŌĆāCommon | 63 (22.7) | 0 (0.0) | 0 (0.0) | ||
| Intralobular inflammation | 309.593 | <0.001 | |||
| ŌĆāNon | 0 (0.0)*,ŌĆĀ | 80 (96.4)ŌĆĀ | 40 (100.0) | ||
| ŌĆā<2 under 20├Ś | 198 (71.5) | 3 (3.6) | 0 (0.0) | ||
| ŌĆā2ŌĆō4 under 20├Ś | 72 (26.0) | 0 (0.0) | 0 (0.0) | ||
| ŌĆā>4 under 20├Ś | 7 (2.5) | 0 (0.0) | 0 (0.0) | ||
| Fibrosis classification | 80.99 | <0.001 | |||
| ŌĆā0 | 94 (34.0)*,ŌĆĀ | 81 (97.6)ŌĆĀ | 40 (100.0) | ||
| ŌĆā1 | 130 (47.0) | 2 (2.4) | 0 (0.0) | ||
| ŌĆā2 | 45 (16.2) | 0 (0.0) | 0 (0.0) | ||
| ŌĆā3 | 7 (2.5) | 0 (0.0) | 0 (0.0) | ||
| ŌĆā4 | 1 (0.3) | 0 (0.0) | 0 (0.0) |
Values are presented as mean┬▒standard deviation, median (interquartile range), or number (%).
P<0.05 was considered to be significant different.
NAFLD, nonalcoholic fatty liver disease; NASH, nonalcoholic steatohepatitis; BMI, body mass index; LCN2, lipocalin-2; HDL, high density lipoprotein; LDL, low density lipoprotein.
Table┬Ā2.
Abbreviations
REFERENCES
- TOOLS
-
METRICS

- ORCID iDs
-
Ming-Hua Zheng

https://orcid.org/0000-0003-4984-2631Jian Wu

https://orcid.org/0000-0001-9933-7364Chunming Ding

https://orcid.org/0000-0002-2063-5594 - Related articles
-
The role of different viral biomarkers on the management of chronic hepatitis B2023 April;29(2)
Novel biomarkers for the management of chronic hepatitis B2020 July;26(3)
Toll-like receptor 9, a possible blocker of non-alcoholic steatohepatitis?2020 April;26(2)
New perspectives of biomarkers for the management of chronic hepatitis B2016 December;22(4)




 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Supplement1
Supplement1 Print
Print